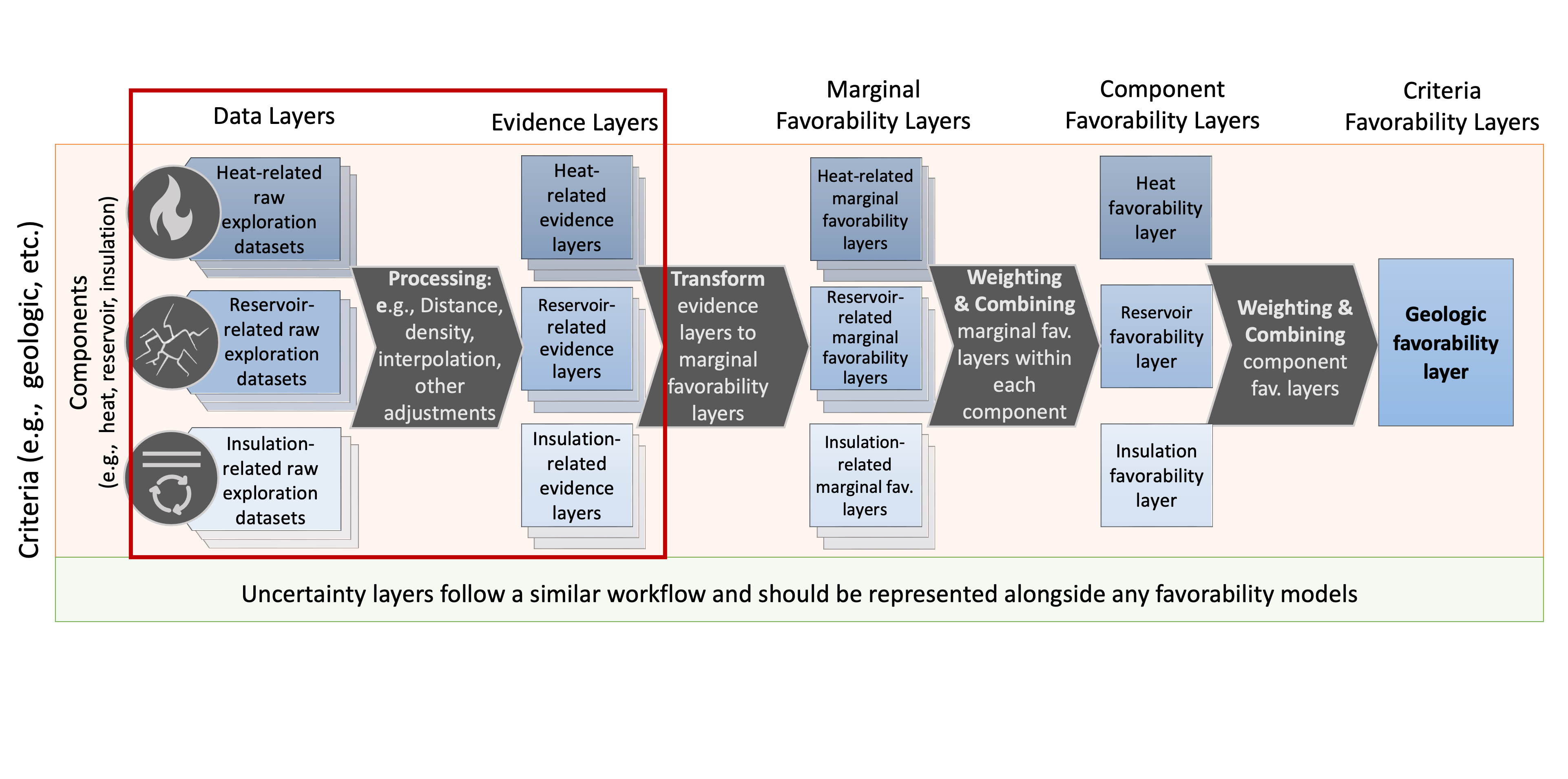
Data Processing with geoPFA: 2D Example from Newberry Volcano, OR¶
geoPFA.Load a PFA configuration file (which defines criteria, components, and layers)
Read raw geospatial datasets into the working
pfadictionaryProcess and clean these datasets into a consistent grid ready for layer combination and favorability modeling
Subsequent notebooks will build upon this by combining layers, applying weights, and producing final favorability models.
1. Imports and Setup¶
# --- General imports ---
from pathlib import Path
import geopandas as gpd
# --- geoPFA core classes ---
from geopfa.data_readers import GeospatialDataReaders
from geopfa.processing import Cleaners, Processing
from geopfa.plotters import GeospatialDataPlotters
# --- Utilities ---
from rex.utilities.utilities import safe_json_load
import json
Define Directories¶
# Get the directory where this notebook lives
notebook_dir = Path(__file__).resolve().parent if "__file__" in locals() else Path.cwd()
# Define the main project directory (where 'config/', 'data/', and 'notebooks/' live)
project_dir = notebook_dir.parent
# Define subdirectories
config_dir = project_dir / "config"
data_dir = project_dir / "data"
# Confirm structure
print("Notebook directory:", notebook_dir)
print("Config directory:", config_dir)
print("Data directory:", data_dir)
# Quick check before proceeding
for folder in [config_dir, data_dir]:
if not folder.exists():
raise FileNotFoundError(f"Expected folder not found: {folder}")
Notebook directory: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/notebooks
Config directory: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/config
Data directory: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data
2. Configuration and Data Input¶
With the project directories defined, the next step is to read the configuration file and load all raw geospatial data into memory.
/data
directory must exactly match those specified in the JSON (see example
below). Folder names must match criteria and component names, and file
names must match layer names.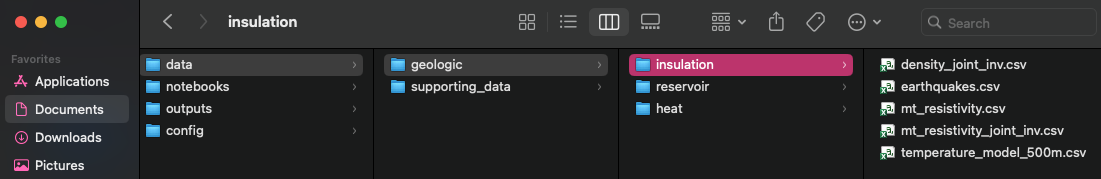
dir_structure¶
Read configuration file¶
# Path to configuration file
pfa_path = config_dir / "newberry_superhot_config.json"
# Load JSON configuration safely (handles comments and malformed JSON)
pfa = safe_json_load(str(pfa_path))
# Quick check
if not pfa_path.exists():
raise FileNotFoundError(f"Configuration file not found: {pfa_path}")
pfa = safe_json_load(str(pfa_path))
print(f"Loaded PFA configuration from: {pfa_path}")
Loaded PFA configuration from: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/config/newberry_superhot_config.json
Load data into the pfa dictionary¶
# Define the file types to be read (others will be skipped with a warning)
file_types = [".csv", ".shp"]
# Gather data according to the config structure
pfa = GeospatialDataReaders.gather_data(data_dir, pfa, file_types)
criteria: geologic
component: heat
reading layer: density_joint_inv
reading layer: mt_resistivity_joint_inv
reading layer: temperature_model_500m
reading layer: earthquakes
reading layer: velocity_model_vs
reading layer: velocity_model_vpvs
component: reservoir
reading layer: lineaments
reading layer: density_joint_inv
reading layer: mt_resistivity_joint_inv
reading layer: earthquakes
reading layer: velocity_model_vs
reading layer: velocity_model_vpvs
component: insulation
reading layer: density_joint_inv
reading layer: mt_resistivity_joint_inv
reading layer: earthquakes
reading layer: temperature_model_500m
reading layer: velocity_model_vp
Verify that all layers loaded correctly¶
This simple diagnostic ensures that each layer entry in the pfa
dictionary contains a "data" key with a valid GeoDataFrame.
missing_data_layers = []
for criteria, crit_data in pfa.get("criteria", {}).items():
for component, comp_data in crit_data.get("components", {}).items():
for layer, layer_data in comp_data.get("layers", {}).items():
if "data" not in layer_data:
missing_data_layers.append({
"criteria": criteria,
"component": component,
"layer": layer,
})
if missing_data_layers:
print("Layers missing 'data':")
for item in missing_data_layers:
print(f" - {item['criteria']} / {item['component']} / {item['layer']}")
else:
print("All layers contain 'data'.")
All layers contain 'data'.
3. Harmonizing Data¶
Now that all raw layers are loaded into the pfa dictionary, we’ll: -
Define a target coordinate reference system (CRS) - Constrain grid
resolution (nx and ny) - Import supporting data such as an area
outline or a well path to give context - Derive a shared spatial extent
for clipping and gridding - Preview the input data layers
Define the Target CRS and Import Contextual Data¶
# EPSG code for UTM Zone 10N (meters)
target_crs = 26910
# Define grid resolution (number of cells in x and y)
nx = 300; ny = 300
# Load the project boundary shapefile
outline_path = data_dir / "supporting_data" / "national_monument_boundary" / "NNVM_bounds.shp"
outline = gpd.read_file(outline_path).to_crs(target_crs)
print(f"Loaded boundary: {outline_path.name}")
Loaded boundary: NNVM_bounds.shp
Unify CRS and Define Extent¶
CRS is defined manually using the variable target_crs defined above.
Extent is defined using the defined extent_layer above, where the
project extent is set to that of the specified extent_layer.
# Reproject all layers to the target CRS
pfa = Cleaners.set_crs(pfa, target_crs=target_crs)
# Choose one representative layer to define the spatial extent
extent_layer = (
pfa["criteria"]["geologic"]["components"]["heat"]["layers"]["mt_resistivity_joint_inv"]["data"]
)
extent = Cleaners.get_extent(extent_layer, dim=2) # Ensure we grab the 2d extent from 3d data
# Validate the extent
print(f"Extent: {extent}")
if extent[0] >= extent[2] or extent[1] >= extent[3]:
raise ValueError("Invalid extent: min values must be less than max values.")
# Define the active criterion (geologic only in this example)
criteria = "geologic"
Extent: [624790.891073, 4825350.71118, 653145.891073, 4855310.71118]
4. Convert 3D Layers to 2D¶
Processing.convert_3d_to_2d() to collapse each 3D
layer along the Z-dimension into a 2D GeoDataFrame.Instead of taking a slice at a given depth, this function sums over Z and each X,Y location, provinding insight into thickness of features. If you’d prefer to use a slice at a given depth, use that 2D slice as your input data layer rather than inputting the 3D model.
"is_3d": "yes"
in the configuration file"needs_flattening": "no")To save time, each unique layer is flattened only once and then reused for any other component that references it. This is especially useful for large shared datasets.
# Cache to store flattened layers and reuse them
flattened_layers = {}
for component, comp_data in pfa["criteria"][criteria]["components"].items():
for layer, layer_config in comp_data["layers"].items():
is_3d = layer_config.get("is_3d", None)
needs_flattening = layer_config.get("needs_flattening", "yes")
# Skip layers not flagged as 3D or marked "no" for flattening
if not is_3d or str(needs_flattening).lower() == "no":
continue
# Use the layer name as the unique cache key
key = layer
if key in flattened_layers:
print(f"Reusing flattened layer: {layer} for component: {component}")
# Copy cached flattened layer into current component/layer
pfa["criteria"][criteria]["components"][component]["layers"][layer] = (
flattened_layers[key].copy()
)
continue
print(f"Converting 3D layer: {layer} → 2D")
# Perform flattening and store in PFA
pfa = Processing.convert_3d_to_2d(pfa, criteria, component=component, layer=layer)
# Cache the flattened result for reuse
flattened_layers[key] = (
pfa["criteria"][criteria]["components"][component]["layers"][layer].copy()
)
print("3D → 2D flattening complete.")
Converting 3D layer: density_joint_inv → 2D
Converting 3D layer: mt_resistivity_joint_inv → 2D
Converting 3D layer: temperature_model_500m → 2D
Converting 3D layer: velocity_model_vs → 2D
Converting 3D layer: velocity_model_vpvs → 2D
Reusing flattened layer: density_joint_inv for component: reservoir
Reusing flattened layer: mt_resistivity_joint_inv for component: reservoir
Reusing flattened layer: velocity_model_vs for component: reservoir
Reusing flattened layer: velocity_model_vpvs for component: reservoir
Reusing flattened layer: density_joint_inv for component: insulation
Reusing flattened layer: mt_resistivity_joint_inv for component: insulation
Reusing flattened layer: temperature_model_500m for component: insulation
Converting 3D layer: velocity_model_vp → 2D
3D → 2D flattening complete.
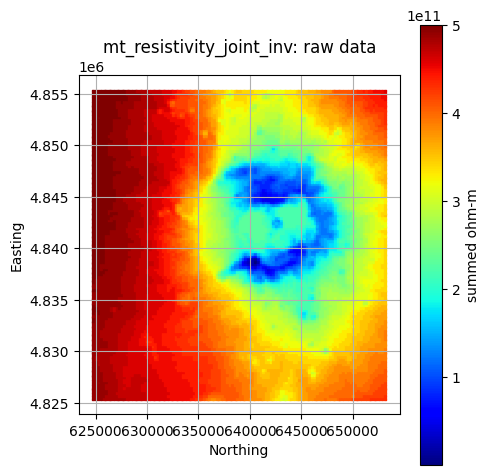
Visualize Cleaned Raw Data Layers¶
geoPFA has built-in plotting tools to help visualize data. Some
examples are shown below using GeospatialDataPlotters.geo_plot_3d.
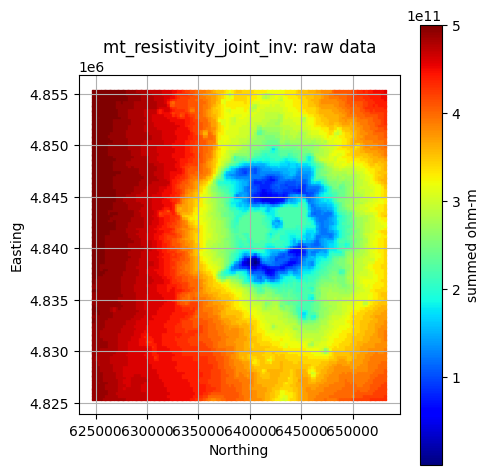
#### Plot a Single Layer
component = "heat"
layer = "mt_resistivity_joint_inv"
gdf = pfa["criteria"][criteria]["components"][component]["layers"][layer]["data"]
col = pfa["criteria"][criteria]["components"][component]["layers"][layer]["data_col"]
units = pfa["criteria"][criteria]["components"][component]["layers"][layer]["units"]
title = f"{layer}: raw data"
GeospatialDataPlotters.geo_plot(gdf, col, units, title, markersize=2, figsize=(5, 5))
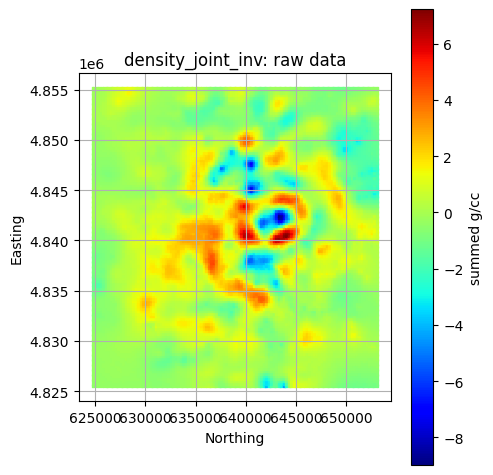
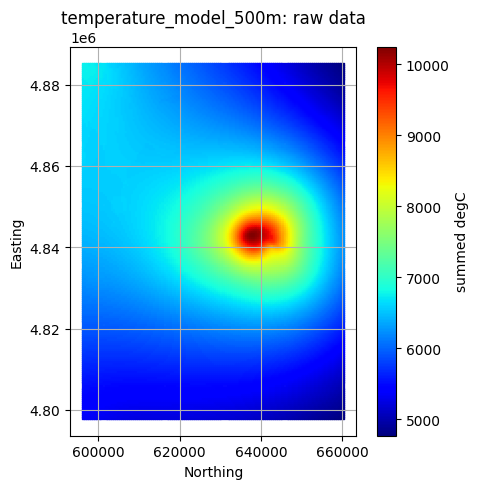
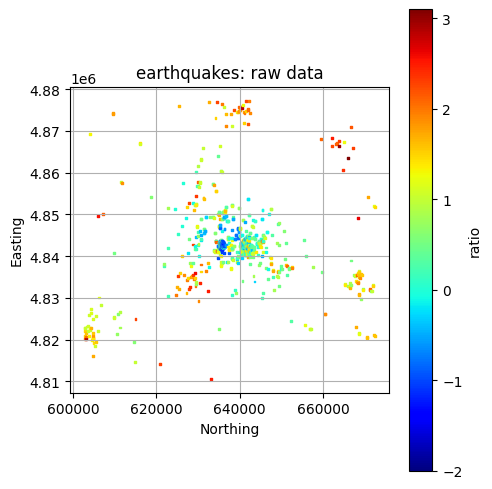
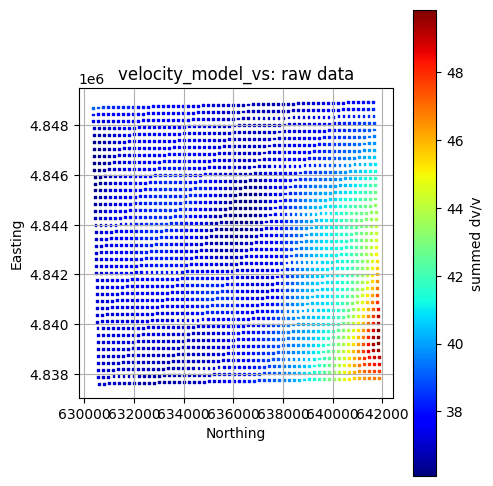
Plot All Layers (Optional)¶
Loop through the full hierarchy and visualize every dataset. This is useful for verifying that CRS, units, and extents align, but can take time for large inputs.
for criteria, crit_data in pfa["criteria"].items():
print(criteria)
plotted_layers = []
for component, comp_data in crit_data["components"].items():
print(f"\t{component}")
for layer, layer_data in comp_data["layers"].items():
if layer in plotted_layers:
print(f"\t\t{layer} plotted already")
continue
gdf = layer_data["data"]
col = layer_data.get("data_col", None)
if layer == "faults_newberry":
col = "None" # faults are geometries only
units = layer_data["units"]
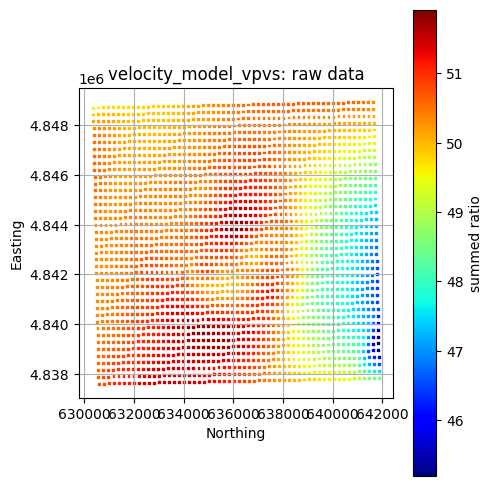
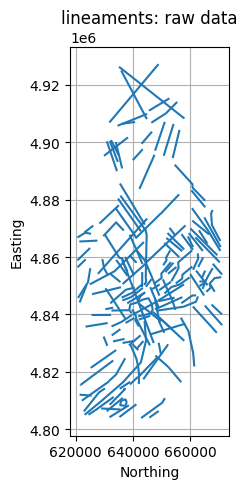
title = f"{layer}: raw data"
GeospatialDataPlotters.geo_plot(gdf, col, units, title, markersize=2, figsize=(5, 5))
plotted_layers.append(layer)
geologic
heat
reservoir
density_joint_inv plotted already
mt_resistivity_joint_inv plotted already
earthquakes plotted already
velocity_model_vs plotted already
velocity_model_vpvs plotted already
insulation
density_joint_inv plotted already
mt_resistivity_joint_inv plotted already
earthquakes plotted already
temperature_model_500m plotted already
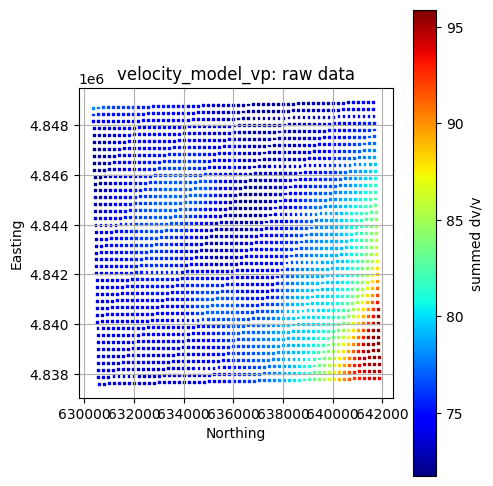
5. Data Processing and Preparation¶
This step transforms raw geospatial inputs into standardized layers on a shared 2D grid for comparison and combination.
(nx, ny) gridHow It Works¶
All 2D geoPFA processing functions follow the same pattern:
Take the raw input stored in
pfa["criteria"][criteria]["components"][component]["layers"][layer]["data"]Process it (e.g., interpolation, point density, distance-to-feature) onto the common 2D grid
Write the result to
pfa["criteria"][criteria]["components"][component]["layers"][layer]["model"]
Filter Outliers¶
Some datasets contain spurious extreme values. Use Cleaners.filter_geodataframe() to remove these outliers based on a quantile threshold.
# Filter the upper 10% of values in selected datasets
target_layers = []
for layer in target_layers:
for component in ["heat", "reservoir"]:
gdf = pfa["criteria"][criteria]["components"]["heat"]["layers"][layer]["data"]
col = pfa["criteria"][criteria]["components"]["heat"]["layers"][layer]["data_col"]
filtered = Cleaners.filter_geodataframe(gdf, col, quantile=0.9)
pfa["criteria"][criteria]["components"][component]["layers"][layer]["data"] = filtered
print("Outlier filtering complete.")
Outlier filtering complete.
Extrapolation¶
geoPFA can perform extrapolation over each layer to ensure every
layer fills the complete extent. geoPFA uses a sparse Gaussian
process regression (GPR) model to backfill missing data. Extrapolation
works best when the extent of the missing data is relatively small
compared to the known data. For large areas, geoPFA reverst to a
linear trend across the training site to produce essentially the average
spatial proess of the known data.
For large datasets, extrapolation can take substanial processing time to
complete. Users can improve model fitting time by reducing the
training_size. Datasets with relatively complete information will
benefit greatly from reduced training sizes and should see little
reduction in performance. An optional verbose flag will enable model
assessment and reporting along with model diagnostic and final
prediction plots.
target_layers = ['velocity_model_vp','velocity_model_vs','velocity_model_vpvs']
extrapolated_layers = {}
for criteria, crit_data in pfa["criteria"].items():
print(criteria)
for component, comp_data in crit_data["components"].items():
print(f"\t{component}")
for layer, layer_config in comp_data["layers"].items():
if layer in extrapolated_layers:
print(f"\t\tReusing cached extrapolation for {layer}")
pfa["criteria"][criteria]["components"][component][
"layers"
][layer] = extrapolated_layers[layer].copy()
elif layer in target_layers:
print(f"\t\tExtrapolating layer: {layer}")
gdf = "data"
col = pfa["criteria"][criteria]["components"][component]["layers"][layer]["data_col"]
pfa = Processing.extrapolate_2d(
pfa,
criteria=criteria,
component=component,
layer=layer,
dataset=gdf,
data_col=col,
training_size=0.8,
verbose=False,
)
extrapolated_layers[layer] = pfa["criteria"][criteria][
"components"
][component]["layers"][layer].copy()
else:
continue
geologic
heat
Extrapolating layer: velocity_model_vs
Extrapolating layer: velocity_model_vpvs
reservoir
Reusing cached extrapolation for velocity_model_vs
Reusing cached extrapolation for velocity_model_vpvs
insulation
Extrapolating layer: velocity_model_vp
Interpolation¶
For most datasets, data can be processed and prepared for layer
combination and favorability estimation by simply interpolating data to
the target grid. Here we do that only once for each input layer that has
"processing_method": "interpolate" specified in the configuration
file.
method = 'linear'
interpolated_layers = {}
for criteria, crit_data in pfa["criteria"].items():
print(criteria)
for component, comp_data in crit_data["components"].items():
print(f"\t{component}")
for layer, layer_config in comp_data["layers"].items():
if layer_config.get("processing_method") == "interpolate":
if layer in interpolated_layers:
print(f"\t\tReusing cached interpolation for {layer}")
pfa["criteria"][criteria]["components"][component]["layers"][layer]['model'] = (
interpolated_layers[layer]['model'].copy()
)
pfa["criteria"][criteria]["components"][component]["layers"][layer]['model_data_col'] = (
interpolated_layers[layer]['model_data_col']
)
pfa["criteria"][criteria]["components"][component]["layers"][layer]['model_units'] = (
interpolated_layers[layer]['model_units']
)
else:
print(f"\t\tInterpolating layer: {layer}")
pfa = Processing.interpolate_points(
pfa,
criteria=criteria,
component=component,
layer=layer,
interp_method=method,
nx=nx,
ny=ny,
extent=extent,
)
interpolated_layers[layer] = (
pfa["criteria"][criteria]["components"][component]["layers"][layer].copy()
)
geologic
heat
Interpolating layer: density_joint_inv
Interpolating layer: mt_resistivity_joint_inv
Interpolating layer: temperature_model_500m
Interpolating layer: velocity_model_vs
Interpolating layer: velocity_model_vpvs
reservoir
Reusing cached interpolation for density_joint_inv
Reusing cached interpolation for mt_resistivity_joint_inv
Reusing cached interpolation for velocity_model_vs
Reusing cached interpolation for velocity_model_vpvs
insulation
Reusing cached interpolation for density_joint_inv
Reusing cached interpolation for mt_resistivity_joint_inv
Reusing cached interpolation for temperature_model_500m
Interpolating layer: velocity_model_vp
Point Density to be Used as a Favorability Proxy¶
For some layers, the spatial density of points (e.g., earthquake
occurrences) is an indicator of permeability or fracturing. geoPFA
has two options to do this in 2D, including: -
Processing.point_density: Computes 2D density projected onto
user-defined grids (ofter coarser than the project grid), ,and then
project those onto the project grid to allow PFA computations that rely
on harmonized grids. - Processing.weighted_distance_from_points:
Compute a weighted influence field from points over the 2D project grid,
using distance to control “influence,” optionallly weighted by an
attribute of the user’s choice (e.g., earthquake magnitude).
In this example, we use Processing.weighted_distance_from_points
because it produces smoother results and alllows us to incorporate
earthquake magnitude. > Tuning Tip: Larger alpha values spread
each equarthquake’s influence farther; smaller values make it more
localized. In other words, a larger value for alpha might be
preferably when wworking with relatively sparse earthquakes.
criteria = 'geologic'
layer = 'earthquakes'
# Filter extreme densities
for comp in ["heat", "reservoir", "insulation"]:
pfa = Processing.weighted_distance_from_points(pfa, criteria=criteria, component=comp,
layer=layer, extent=extent, nx=nx, ny=ny, alpha=1000)
gdf = pfa["criteria"][criteria]["components"][comp]["layers"][layer]["model"].copy()
col = pfa["criteria"][criteria]["components"][comp]["layers"][layer]["model_data_col"]
filtered = Cleaners.filter_geodataframe(gdf, col, quantile=0.9)
pfa["criteria"][criteria]["components"][comp]["layers"][layer]["model"] = filtered
Process Faults¶
process_faults() function to calculate a
fault-based favorability score that increases with proximity to
faults and (optionally) their intersections.alpha_fault, alpha_intersection) to represent how
influence decreases with distance.weight_fault and weight_intersection.Tuning Tip: Larger
alphavalues spread the fault’s influence farther; smaller values make it more localized.
component = "reservoir"
layer = "lineaments"
# --- Fault processing for reservoir component ---
pfa = Processing.process_faults(
pfa,
criteria=criteria,
component=component,
layer=layer,
extent=extent,
nx=nx,
ny=ny,
alpha_fault=5500.0, # decay distance for faults
alpha_intersection=3500.0, # decay distance for intersections
weight_fault=0.7, # relative weighting of faults vs intersections
weight_intersection=0.3,
use_intersections=True, # include intersection effects
)
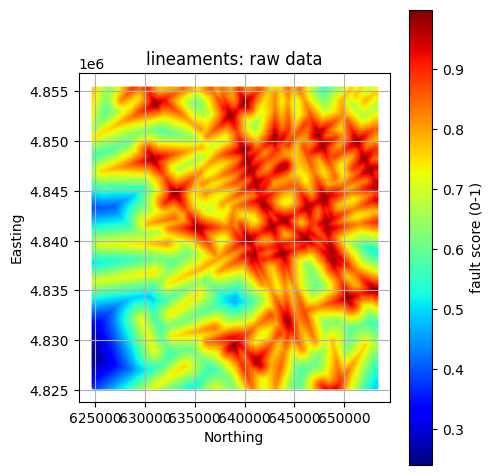
# Visualize the results
# Note the plotted extent differs from the raw data when comparing
# Note we now grab the processed data from its "model" locations
gdf = pfa["criteria"][criteria]["components"][component]["layers"][layer]["model"]
col = pfa["criteria"][criteria]["components"][component]["layers"][layer]["model_data_col"]
units = pfa["criteria"][criteria]["components"][component]["layers"][layer]["model_units"]
title = f"{layer}: raw data"
GeospatialDataPlotters.geo_plot(gdf, col, units, title, markersize=2, figsize=(5, 5))
Plot All Processed Layers¶
for criteria, crit_data in pfa["criteria"].items():
print(criteria)
plotted_layers = []
for component, comp_data in crit_data["components"].items():
print(f"\t{component}")
for layer, layer_data in comp_data["layers"].items():
if layer in plotted_layers:
continue
# Skip if no processed model exists
if "model" not in layer_data or layer_data["model"] is None:
print(f"\t\t!! Skipping {layer}: no processed 'model' found.")
continue
gdf = layer_data["model"]
col = layer_data.get("model_data_col")
units = layer_data.get("model_units", "None")
title = f"{layer}: processed model"
# Plot in 3D
GeospatialDataPlotters.geo_plot(
gdf, col, units, title, area_outline=outline, extent=extent, markersize=0.75, figsize=(5, 5)
)
plotted_layers.append(layer)
print("\nFinished plotting all available processed layers.")
geologic
heat
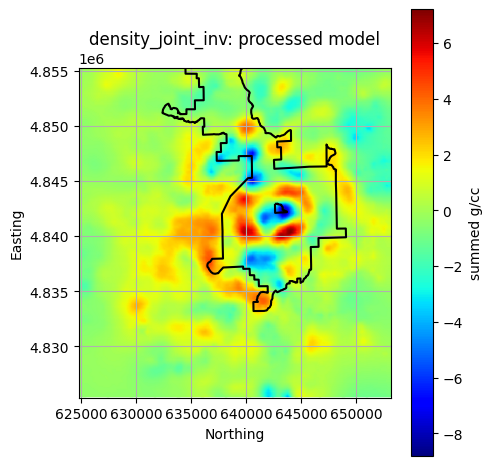
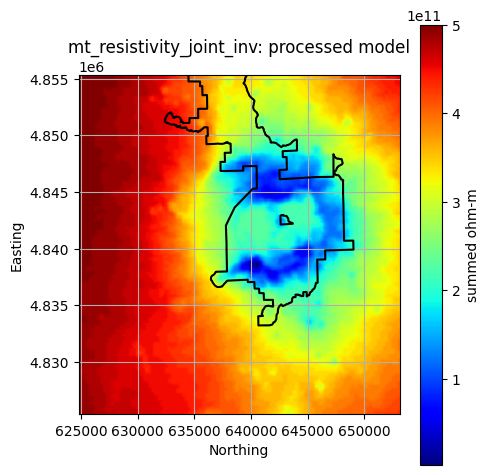
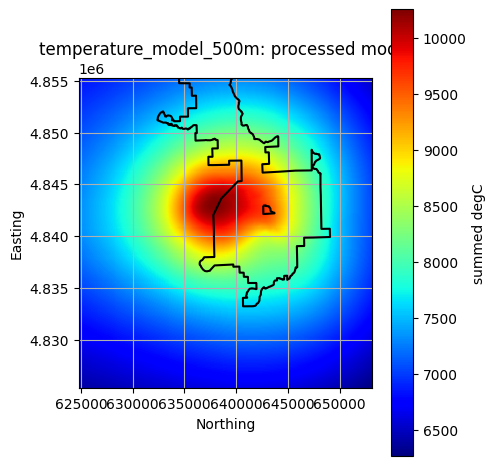
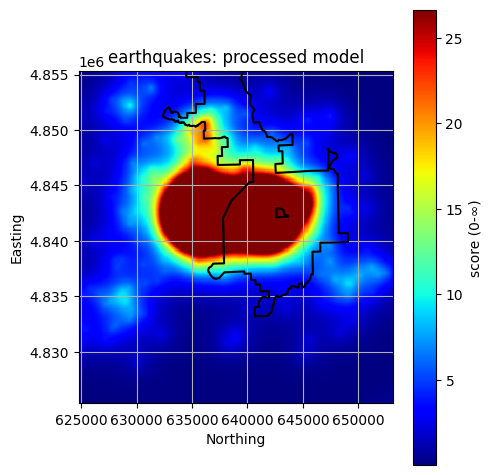
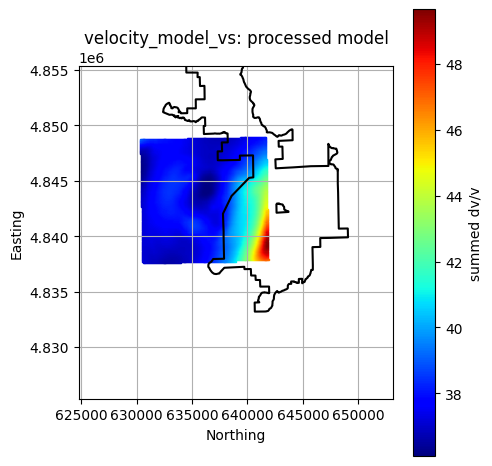
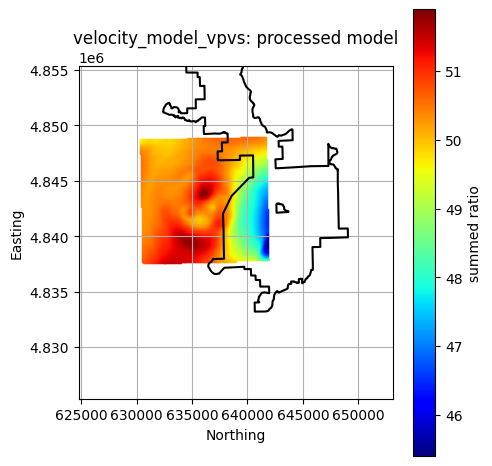
reservoir
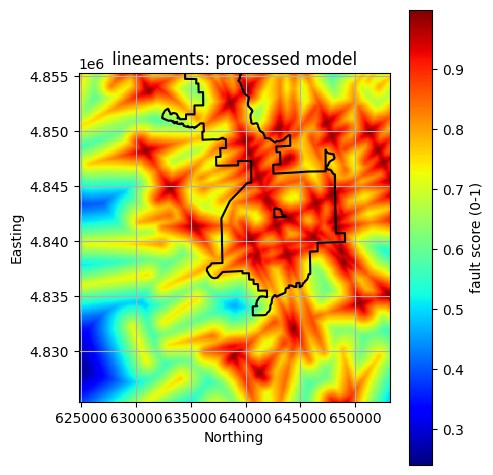
insulation
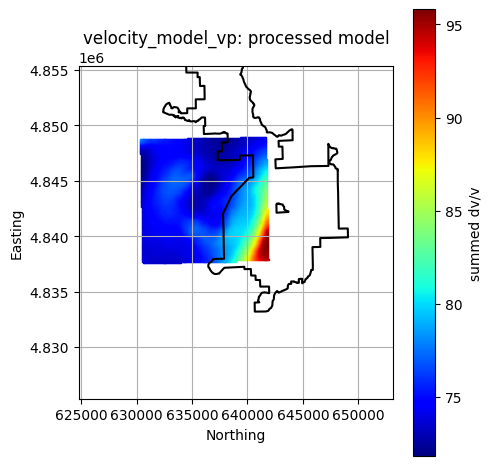
Finished plotting all available processed layers.
Save Processed Layers¶
It’s good practice to save processed data to disk before proceeding so processing only has to be completed once. This allows the user to make minor tweaks to the PFA, such as adjusting weights, without reprocessing all data layers.
for criteria, crit_data in pfa["criteria"].items():
print(criteria)
for component, comp_data in crit_data["components"].items():
print(f"\t{component}")
for layer, layer_data in comp_data["layers"].items():
gdf = layer_data["model"]
col = layer_data["model_data_col"]
units = layer_data["model_units"]
out_fp = data_dir / criteria / component / f"{layer}_processed.csv"
gdf.to_csv(out_fp, index=False)
print(f"\t\tSaved: {out_fp}")
geologic
heat
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/density_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/mt_resistivity_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/temperature_model_500m_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/earthquakes_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/velocity_model_vs_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/heat/velocity_model_vpvs_processed.csv
reservoir
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/density_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/mt_resistivity_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/earthquakes_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/lineaments_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/velocity_model_vs_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/reservoir/velocity_model_vpvs_processed.csv
insulation
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/insulation/density_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/insulation/mt_resistivity_joint_inv_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/insulation/earthquakes_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/insulation/temperature_model_500m_processed.csv
Saved: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/data/geologic/insulation/velocity_model_vp_processed.csv
Save a “Clean” PFA Config (no GeoDataFrames)¶
Similarly, saving a “clean” processed PFA config allows the user to avoid repeating the processing step when making minor adjustments to the PFA. This processed config serves as the config file for the next step.
def drop_geodataframes(data: dict) -> dict:
"""Recursively remove GeoDataFrame objects before saving."""
clean = {}
for key, val in data.items():
if isinstance(val, gpd.GeoDataFrame):
continue
elif isinstance(val, dict):
clean[key] = drop_geodataframes(val)
else:
clean[key] = val
return clean
pfa_nodf = drop_geodataframes(pfa)
out_json = config_dir / "newberry_superhot_processed_config.json"
with open(out_json, "w") as f:
json.dump(pfa_nodf, f, indent=4)
print(f"Processed PFA configuration saved to: {out_json}")
Processed PFA configuration saved to: /Users/ntaverna/Documents/DEEPEN/geoPFA/examples/Newberry/2D/config/newberry_superhot_processed_config.json
Next Steps: Layer Combination and Favorability Modeling¶
This concludes Notebook 1 – Data Processing with ``geoPFA``.
You’re now ready to combine these layers into component and criteria favorability models.